-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
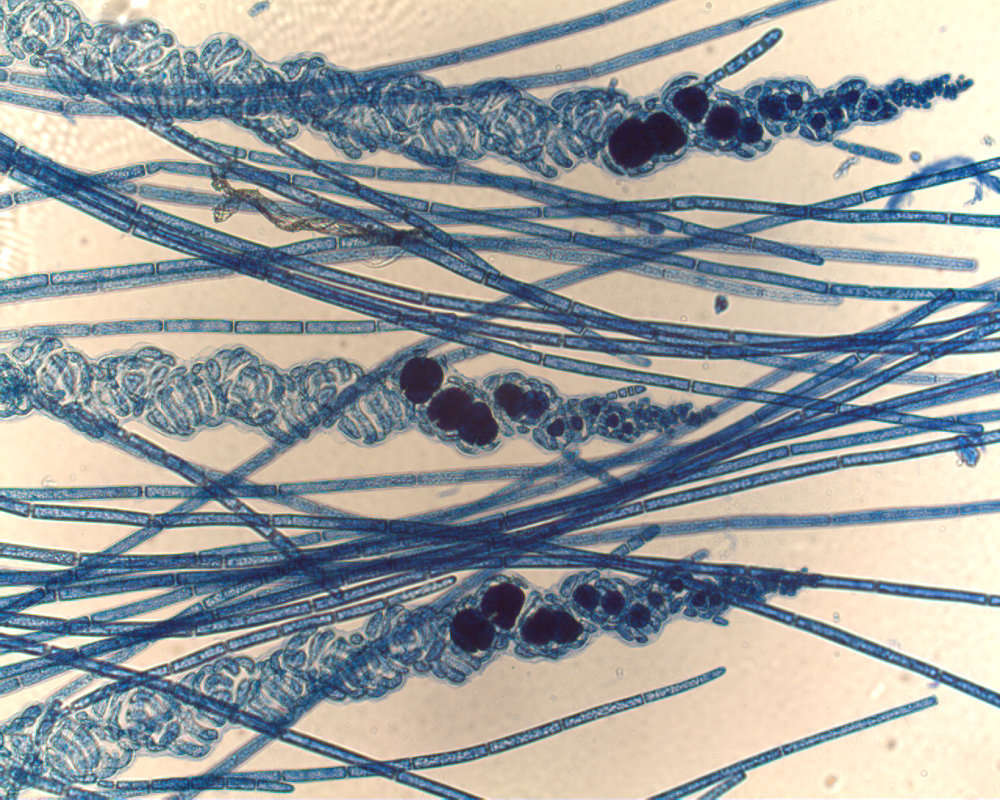
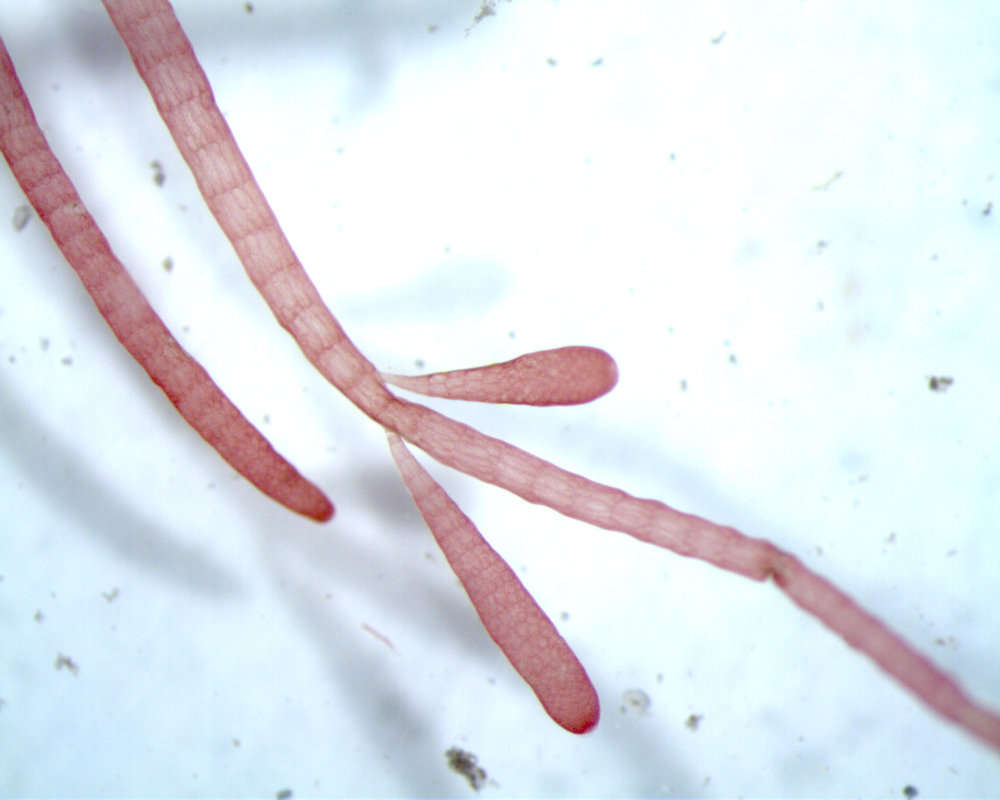
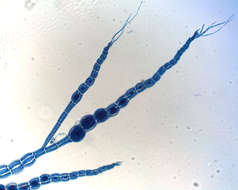
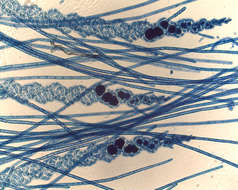
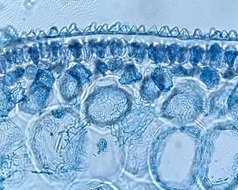
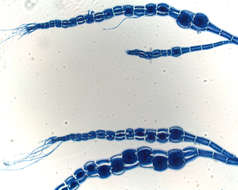
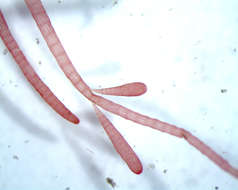
Thallus of Ceramium diaphanum with tetrasporangia. Collected from Bodden, the brackish waters lying between the isles of Hiddensee and Ruegen (German Baltic Sea). This image was taken using Zeiss Universal with Olympus C7070 CCD camera.
-
Fornæs, Djursland, Danmark
-
Fornæs, Djursland, Danmark
-
Fornæs, Djursland, Danmark
-
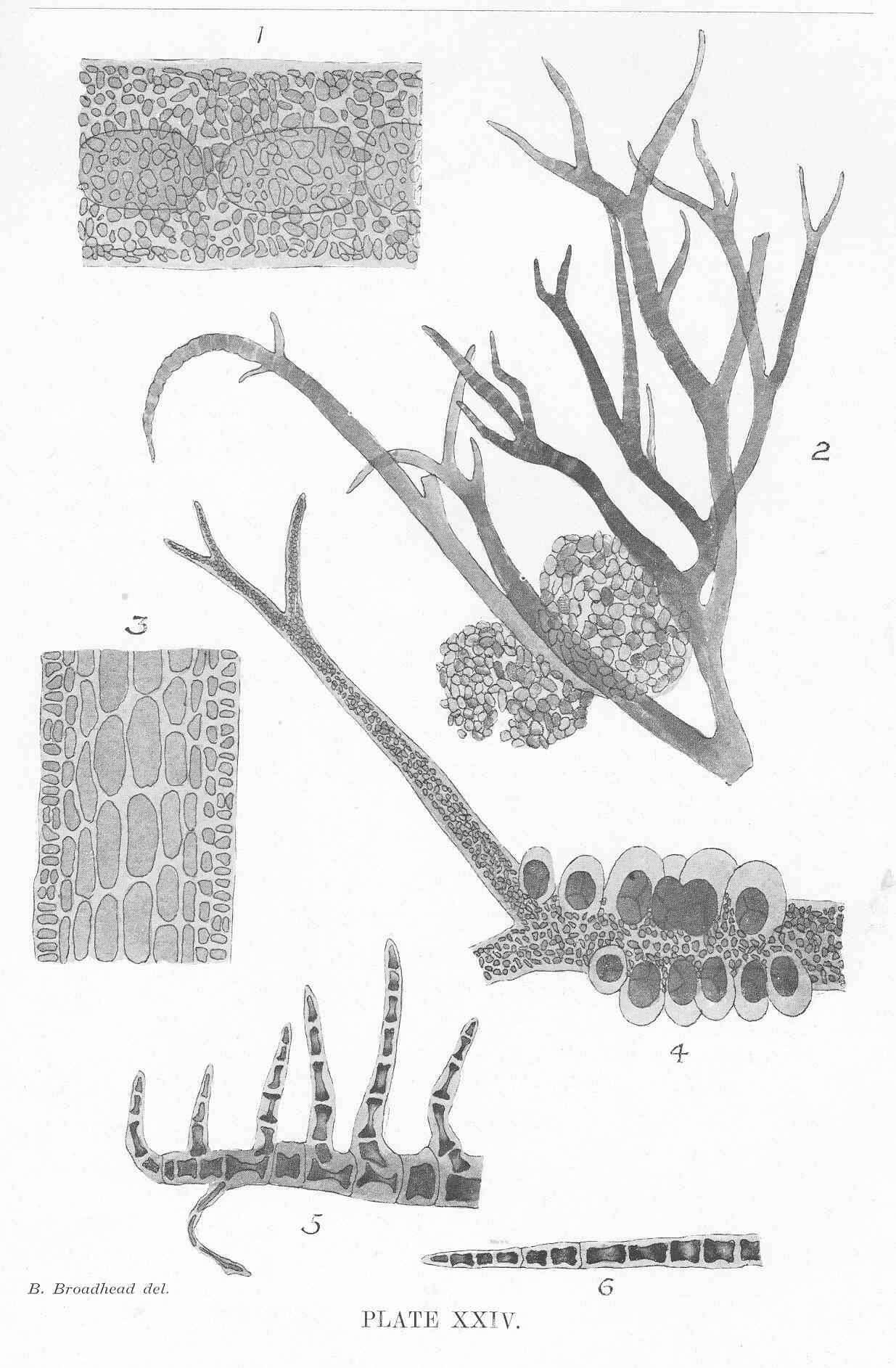
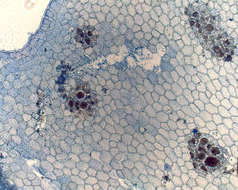
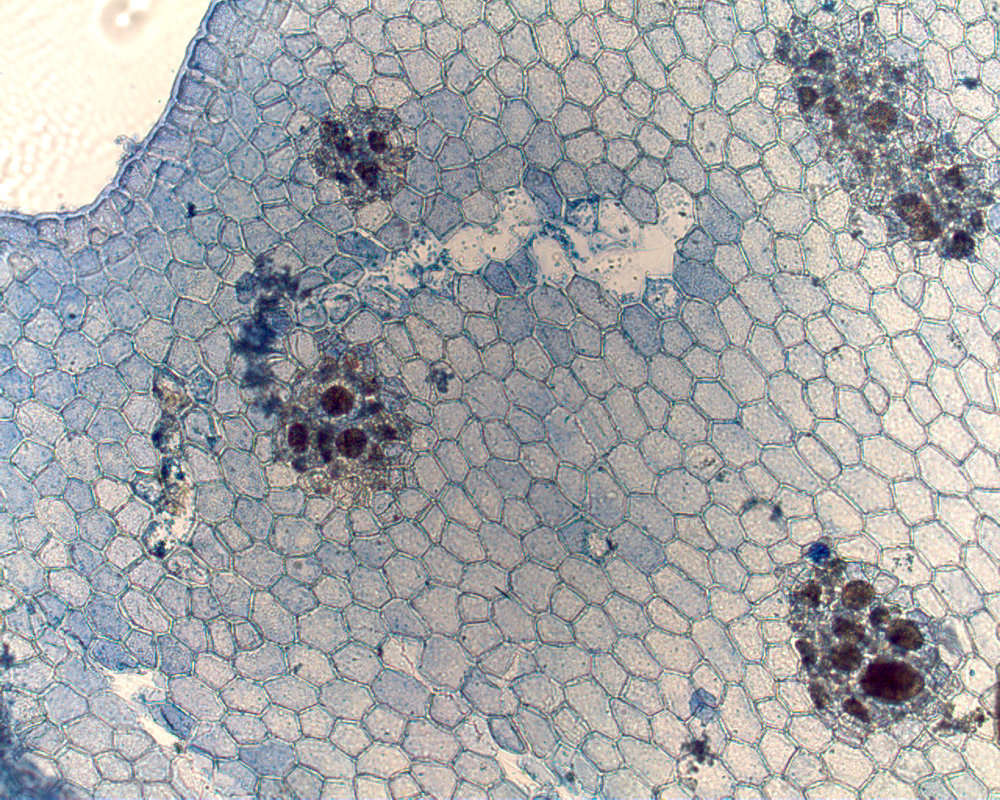
Nitophyllum medium, type, veins evident, tetrasporangial sori in rather irregular lines.
-
Ceramium stichidiosum, Rhodymenia, Rhizoclonium hookeri.
-
Laurencia tuberculosa var. gemmifera.
-
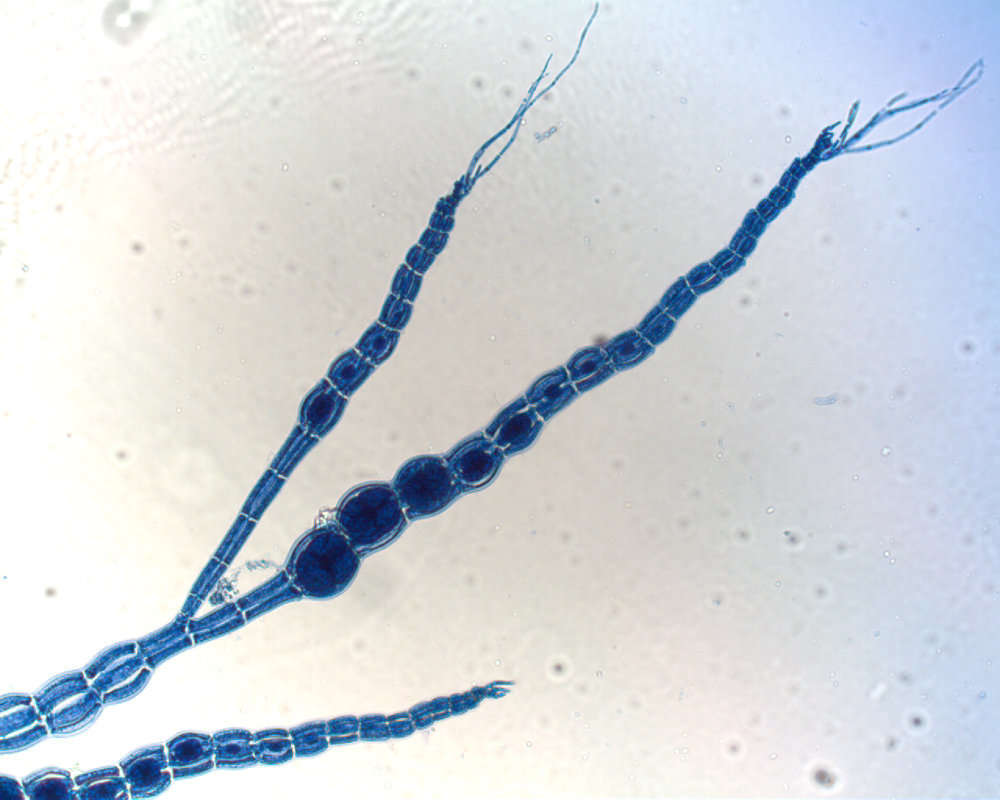
Rhodophycees ou Floridees (Algues rouges) Ceramiees, Griffithsia corallina (Lightf.) Ag..
-
Polysiphonia dendroidea, a piece magnified.
-
Rhodophycees ou Floridees (Algues rouges) Rhodomelees, Brongniartella byssoides (Good et Wood.) Schm..
-
Rhodophycees ou Floridees (Algues rouges) Rhodomelees, Laurencia pinnatifida (Gmel.) Lamour.
-
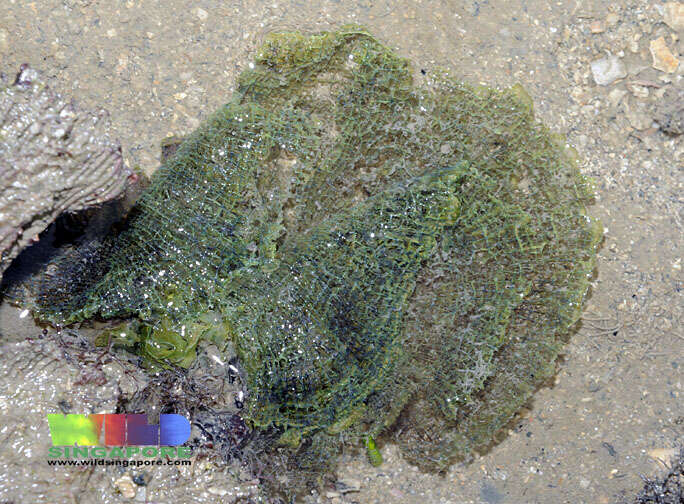
Singapore, South West, Singapore
-
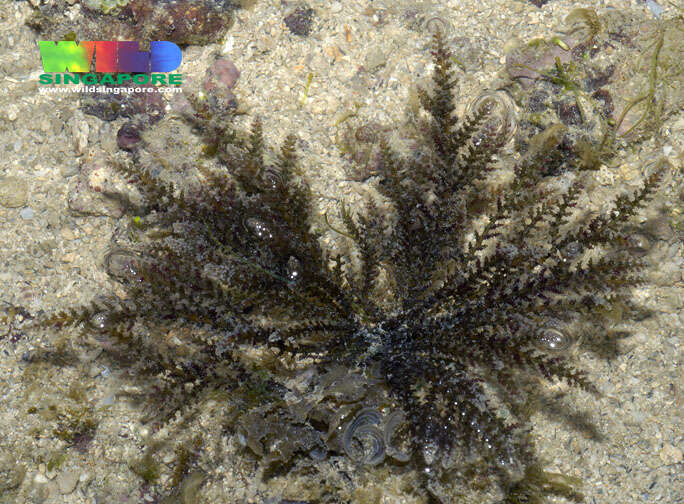
Singapore, Central Singapore, Singapore
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.